Cascade, Selection, Reverse-Engineering and Prediction in Cascade Networks
Frédéric Bertrand and Myriam Maumy-Bertrand
https://doi.org/10.32614/CRAN.package.Cascade
Cascade is a modeling tool allowing gene selection, reverse engineering, and prediction in cascade networks. Jung, N., Bertrand, F., Bahram, S., Vallat, L., and Maumy-Bertrand, M. (2014) https://doi.org/10.1093/bioinformatics/btt705.
The package was presented at the User2014! conference. Jung, N., Bertrand, F., Bahram, S., Vallat, L., and Maumy-Bertrand, M. (2014). “Cascade: a R-package to study, predict and simulate the diffusion of a signal through a temporal genenetwork”, book of abstracts, User2014!, Los Angeles, page 153, https://user2014.r-project.org/abstracts/posters/181_Jung.pdf.
This website and these examples were created by F. Bertrand and M. Maumy-Bertrand.
Installation
You can install the released version of Cascade from CRAN with:
install.packages("Cascade")You can install the development version of Cascade from github with:
devtools::install_github("fbertran/Cascade")Examples
Data management
Import Cascade Data (repeated measurements on several subjects) from the CascadeData package and turn them into a micro array object. The second line makes sure the CascadeData package is installed.
library(Cascade)
if(!require(CascadeData)){install.packages("CascadeData")}
#> Loading required package: CascadeData
data(micro_US)
micro_US<-as.micro_array(micro_US,time=c(60,90,210,390),subject=6)Get a summay and plots of the data:
summary(micro_US)
#> Loading required package: cluster
#> N1_US_T60 N1_US_T90 N1_US_T210 N1_US_T390 N2_US_T60
#> Min. : 1.0 Min. : 1.0 Min. : 1.0 Min. : 1.0 Min. : 1.0
#> 1st Qu.: 19.7 1st Qu.: 18.8 1st Qu.: 15.2 1st Qu.: 20.9 1st Qu.: 18.5
#> Median : 38.0 Median : 37.2 Median : 34.9 Median : 40.2 Median : 36.9
#> Mean : 107.5 Mean : 106.9 Mean : 109.6 Mean : 105.7 Mean : 110.6
#> 3rd Qu.: 80.6 3rd Qu.: 82.1 3rd Qu.: 82.8 3rd Qu.: 84.8 3rd Qu.: 85.3
#> Max. :8587.9 Max. :8311.7 Max. :7930.3 Max. :7841.8 Max. :7750.3
#> N2_US_T90 N2_US_T210 N2_US_T390 N3_US_T60 N3_US_T90
#> Min. : 1.0 Min. : 1.0 Min. : 1.0 Min. : 1.0 Min. : 1.0
#> 1st Qu.: 17.1 1st Qu.: 15.8 1st Qu.: 17.7 1st Qu.: 17.3 1st Qu.: 19.5
#> Median : 36.7 Median : 36.0 Median : 37.4 Median : 34.4 Median : 38.2
#> Mean : 102.1 Mean : 106.8 Mean : 111.3 Mean : 101.6 Mean : 107.1
#> 3rd Qu.: 78.2 3rd Qu.: 83.5 3rd Qu.: 86.4 3rd Qu.: 75.4 3rd Qu.: 82.3
#> Max. :8014.3 Max. :8028.6 Max. :7498.4 Max. :8072.2 Max. :7889.2
#> N3_US_T210 N3_US_T390 N4_US_T60 N4_US_T90 N4_US_T210
#> Min. : 1.0 Min. : 1.0 Min. : 1.0 Min. : 1.0 Min. : 1.0
#> 1st Qu.: 16.4 1st Qu.: 20.9 1st Qu.: 20.4 1st Qu.: 19.5 1st Qu.: 20.5
#> Median : 34.7 Median : 41.0 Median : 38.9 Median : 38.5 Median : 39.9
#> Mean : 100.3 Mean : 113.9 Mean : 113.6 Mean : 114.8 Mean : 110.1
#> 3rd Qu.: 76.3 3rd Qu.: 89.2 3rd Qu.: 84.6 3rd Qu.: 86.1 3rd Qu.: 86.8
#> Max. :8278.2 Max. :6856.2 Max. :9502.3 Max. :9193.4 Max. :9436.0
#> N4_US_T390 N5_US_T60 N5_US_T90 N5_US_T210 N5_US_T390
#> Min. : 1.0 Min. : 1.0 Min. : 1.0 Min. : 1.0 Min. : 1.0
#> 1st Qu.: 19.9 1st Qu.: 16.8 1st Qu.: 18.8 1st Qu.: 19.5 1st Qu.: 19.9
#> Median : 38.8 Median : 34.5 Median : 36.9 Median : 38.2 Median : 39.0
#> Mean : 111.7 Mean : 111.3 Mean : 108.0 Mean : 107.4 Mean : 109.8
#> 3rd Qu.: 85.4 3rd Qu.: 82.0 3rd Qu.: 81.4 3rd Qu.: 82.4 3rd Qu.: 84.9
#> Max. :8771.0 Max. :8569.3 Max. :7970.1 Max. :8371.0 Max. :7686.5
#> N6_US_T60 N6_US_T90 N6_US_T210 N6_US_T390
#> Min. : 1.0 Min. : 1.0 Min. : 1.0 Min. : 1.0
#> 1st Qu.: 21.1 1st Qu.: 21.5 1st Qu.: 19.9 1st Qu.: 20.2
#> Median : 40.9 Median : 40.8 Median : 39.1 Median : 39.4
#> Mean : 110.1 Mean : 108.5 Mean : 112.0 Mean : 109.5
#> 3rd Qu.: 86.3 3rd Qu.: 85.6 3rd Qu.: 86.3 3rd Qu.: 86.6
#> Max. :8241.0 Max. :8355.0 Max. :8207.1 Max. :9520.0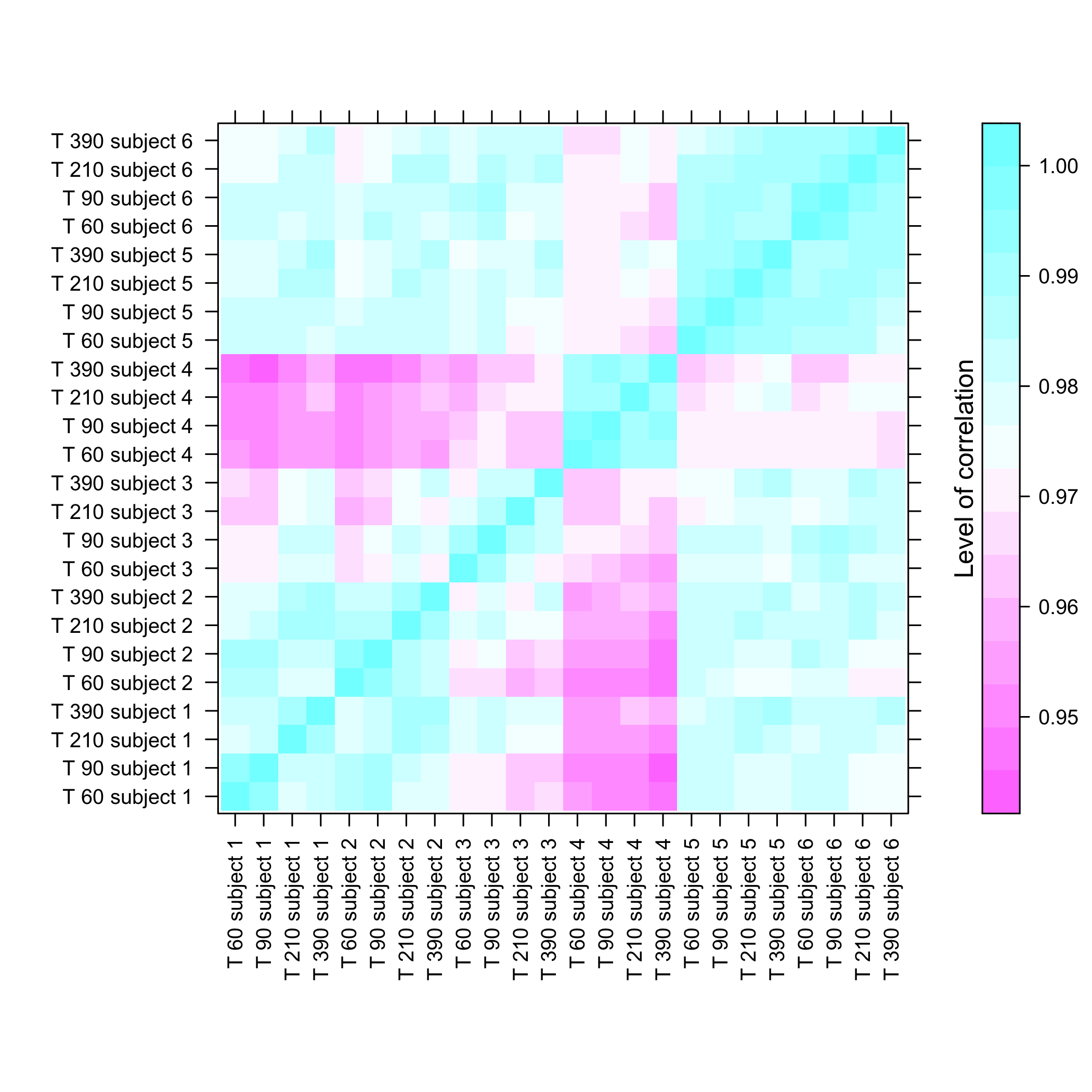
plot of chunk plotmicroarrayclass
plot of chunk plotmicroarrayclass
Gene selection
There are several functions to carry out gene selection before the inference. They are detailed in the package vignette and in the website-only E-MTAB-1475 re-analysis PDF.
Data simulation
Let’s simulate some cascade data and then do some reverse engineering.
We first design the F matrix
T<-4
F<-array(0,c(T-1,T-1,T*(T-1)/2))
for(i in 1:(T*(T-1)/2)){diag(F[,,i])<-1}
F[,,2]<-F[,,2]*0.2
F[2,1,2]<-1
F[3,2,2]<-1
F[,,4]<-F[,,2]*0.3
F[3,1,4]<-1
F[,,5]<-F[,,2]We set the seed to make the results reproducible and draw a scale free random network.
set.seed(1)
Net<-Cascade::network_random(
nb=100,
time_label=rep(1:4,each=25),
exp=1,
init=1,
regul=round(rexp(100,1))+1,
min_expr=0.1,
max_expr=2,
casc.level=0.4
)
Net@F<-FWe simulate gene expression according to the network that was previously drawn
M <- Cascade::gene_expr_simulation(
network=Net,
time_label=rep(1:4,each=25),
subject=5,
level_peak=200)
#> Loading required package: VGAM
#> Loading required package: stats4
#> Loading required package: splinesGet a summay and plots of the simulated data:
summary(M)
#> log(S/US) : P1T1 log(S/US) : P1T2 log(S/US) : P1T3 log(S/US) : P1T4
#> Min. :-887.428 Min. :-2060.695 Min. :-837.5811 Min. :-2189.33
#> 1st Qu.: -53.644 1st Qu.: -70.993 1st Qu.: -85.2176 1st Qu.: -115.23
#> Median : 6.122 Median : 2.206 Median : -6.6303 Median : -10.49
#> Mean : 3.928 Mean : -3.800 Mean : 0.5563 Mean : -75.60
#> 3rd Qu.: 74.894 3rd Qu.: 79.999 3rd Qu.: 71.9501 3rd Qu.: 43.85
#> Max. : 747.779 Max. : 1461.163 Max. :1279.6850 Max. : 492.80
#> log(S/US) : P2T1 log(S/US) : P2T2 log(S/US) : P2T3 log(S/US) : P2T4 log(S/US) : P3T1
#> Min. :-431.905 Min. :-845.072 Min. :-557.840 Min. :-660.17 Min. :-868.834
#> 1st Qu.: -50.282 1st Qu.: -21.668 1st Qu.: -42.958 1st Qu.: -63.34 1st Qu.: -44.656
#> Median : -3.782 Median : 2.059 Median : -2.448 Median : -10.97 Median : 1.839
#> Mean : 8.264 Mean : 18.235 Mean : 25.290 Mean : 18.10 Mean : -2.150
#> 3rd Qu.: 40.975 3rd Qu.: 40.323 3rd Qu.: 46.573 3rd Qu.: 42.56 3rd Qu.: 55.072
#> Max. :1287.819 Max. : 699.912 Max. :1754.081 Max. :1418.99 Max. : 597.562
#> log(S/US) : P3T2 log(S/US) : P3T3 log(S/US) : P3T4 log(S/US) : P4T1
#> Min. :-953.894 Min. :-1182.316 Min. :-1027.24 Min. :-1012.76
#> 1st Qu.: -59.964 1st Qu.: -87.170 1st Qu.: -61.31 1st Qu.: -33.79
#> Median : -1.306 Median : -2.614 Median : 16.27 Median : 11.57
#> Mean : 24.123 Mean : 10.224 Mean : 27.52 Mean : 10.98
#> 3rd Qu.: 78.430 3rd Qu.: 79.246 3rd Qu.: 60.93 3rd Qu.: 74.09
#> Max. :1808.233 Max. : 2761.291 Max. : 1926.45 Max. : 891.60
#> log(S/US) : P4T2 log(S/US) : P4T3 log(S/US) : P4T4 log(S/US) : P5T1
#> Min. :-1569.32 Min. :-577.799 Min. :-661.083 Min. :-555.708
#> 1st Qu.: -118.83 1st Qu.: -62.623 1st Qu.: -41.705 1st Qu.: -64.469
#> Median : -13.84 Median : -8.788 Median : -1.468 Median : 2.697
#> Mean : -63.39 Mean : -14.803 Mean : 10.111 Mean : 9.403
#> 3rd Qu.: 40.22 3rd Qu.: 37.779 3rd Qu.: 67.888 3rd Qu.: 57.251
#> Max. : 678.27 Max. : 430.737 Max. : 492.723 Max. : 654.771
#> log(S/US) : P5T2 log(S/US) : P5T3 log(S/US) : P5T4
#> Min. :-1467.268 Min. :-911.18 Min. :-621.705
#> 1st Qu.: -68.769 1st Qu.: -71.66 1st Qu.: -62.466
#> Median : -1.565 Median : 1.54 Median : -5.789
#> Mean : 8.180 Mean : 13.03 Mean : 7.083
#> 3rd Qu.: 62.256 3rd Qu.: 86.29 3rd Qu.: 62.041
#> Max. : 990.550 Max. :1386.11 Max. : 689.323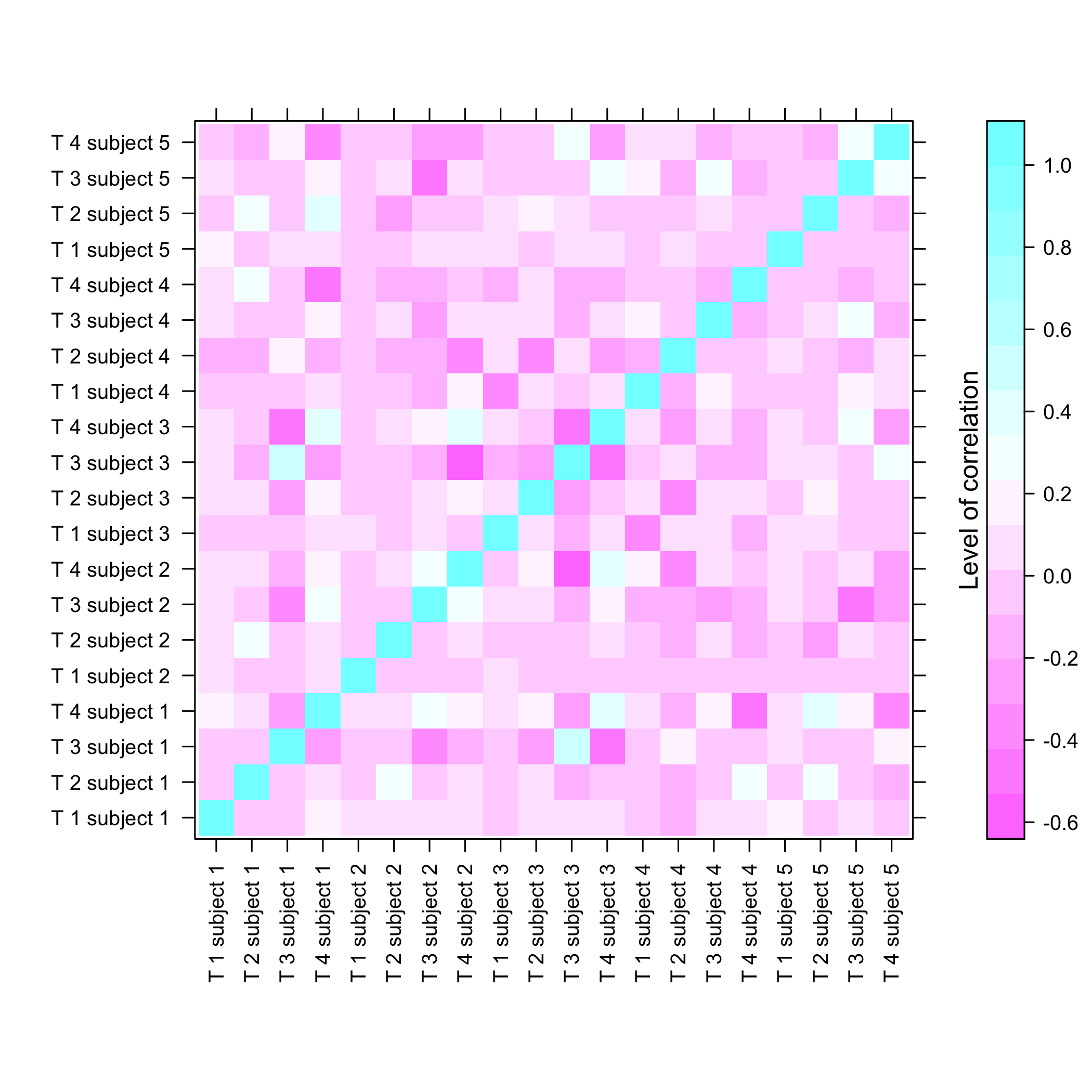
plot of chunk summarysimuldata
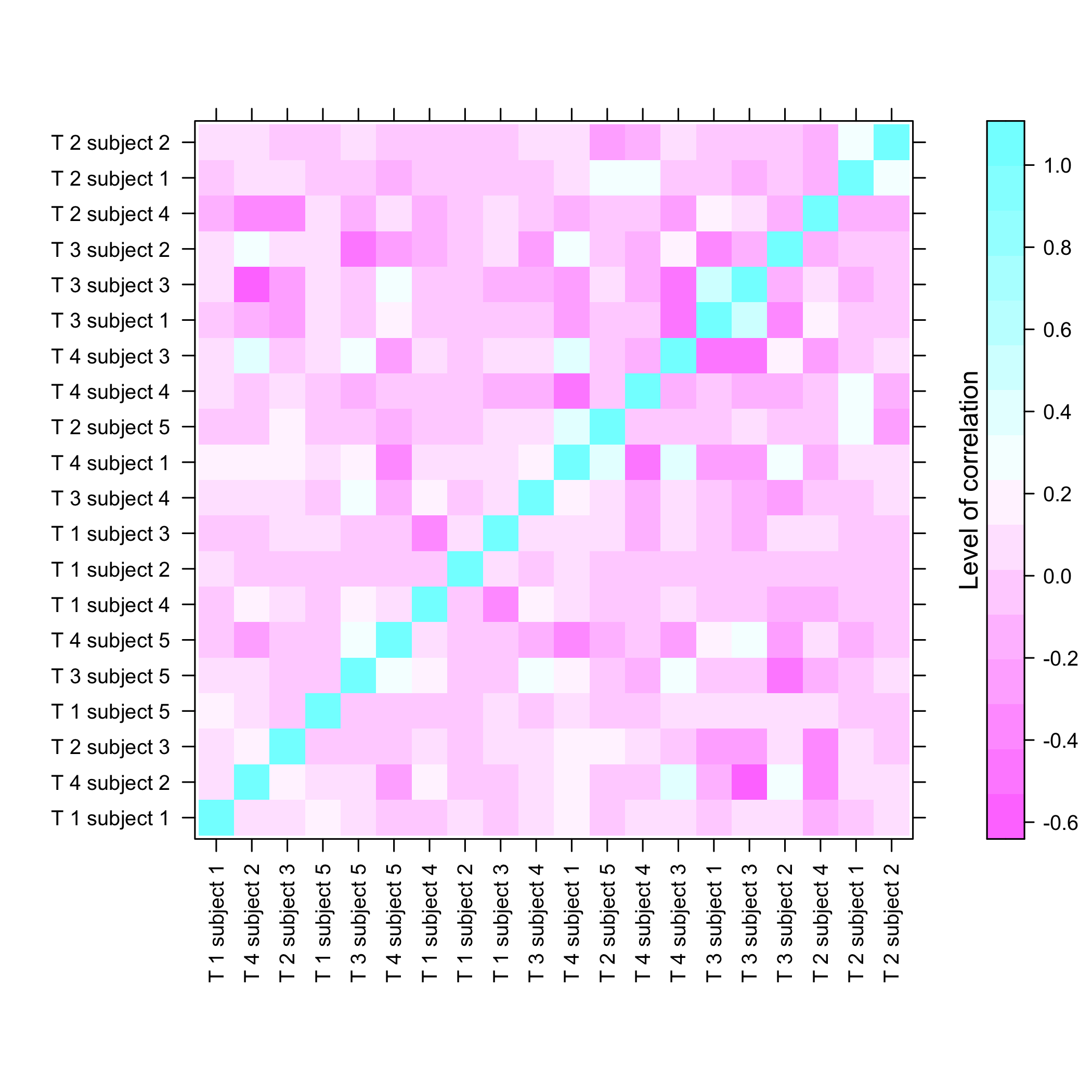
plot of chunk summarysimuldata
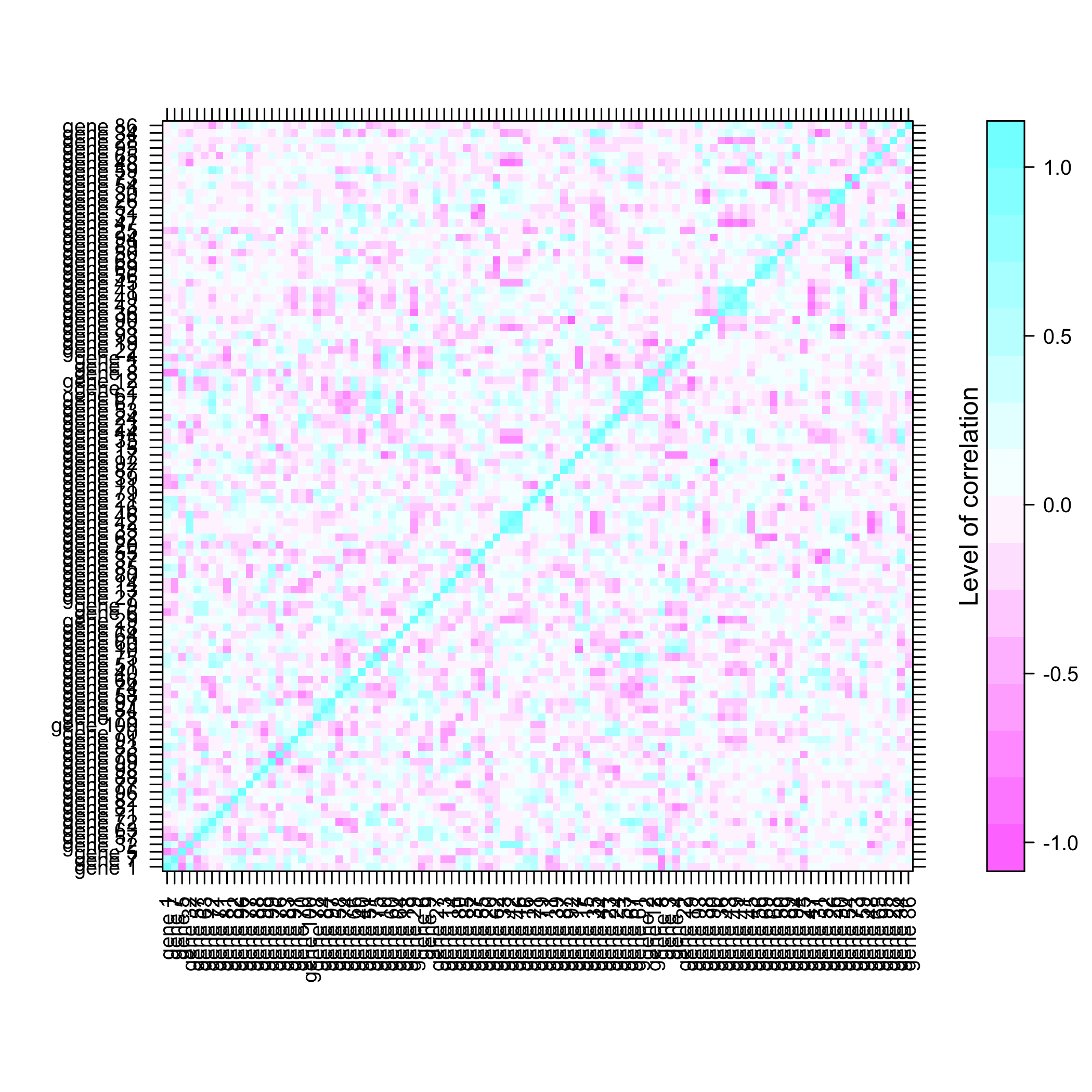
plot of chunk summarysimuldata
plot(M)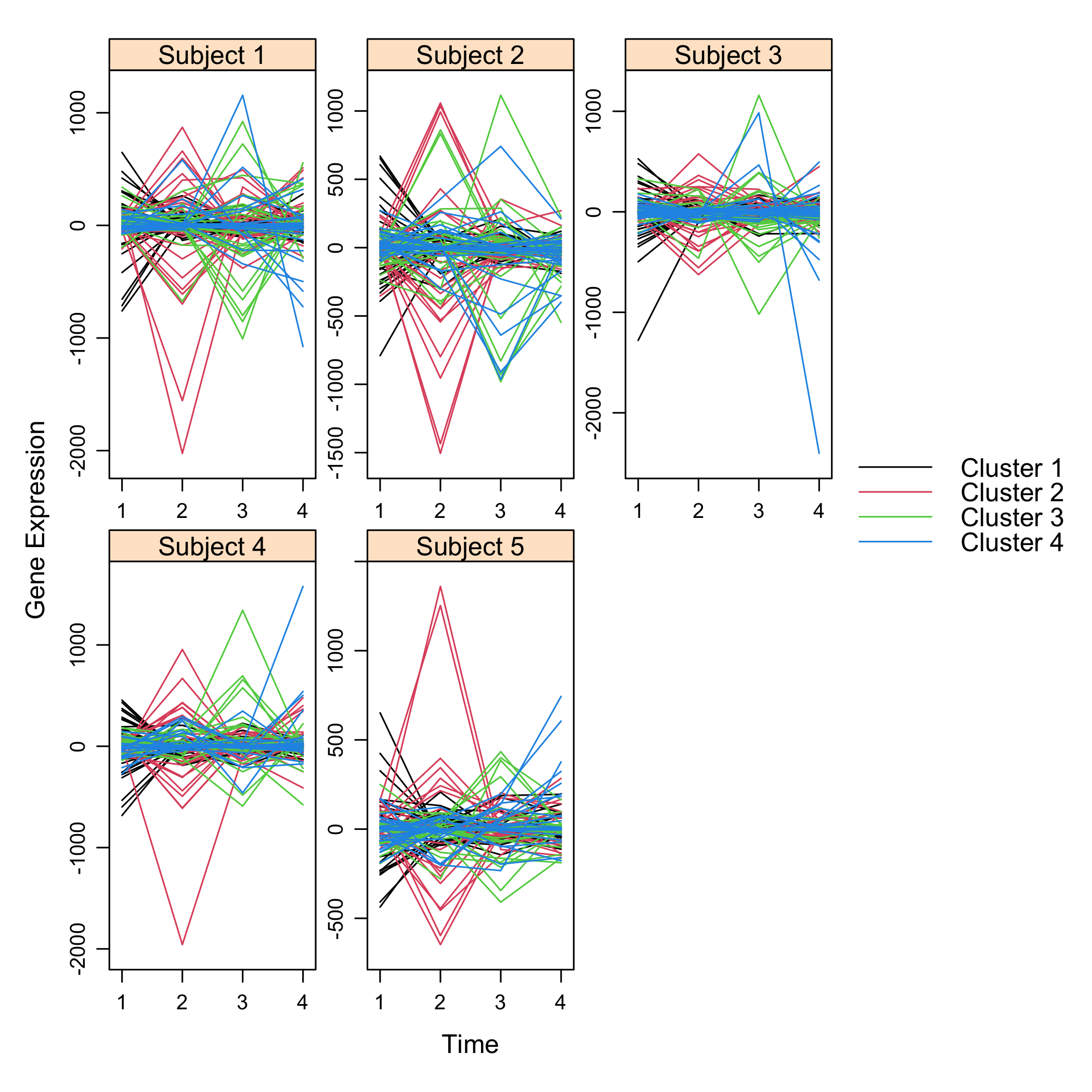
plot of chunk plotsimuldata
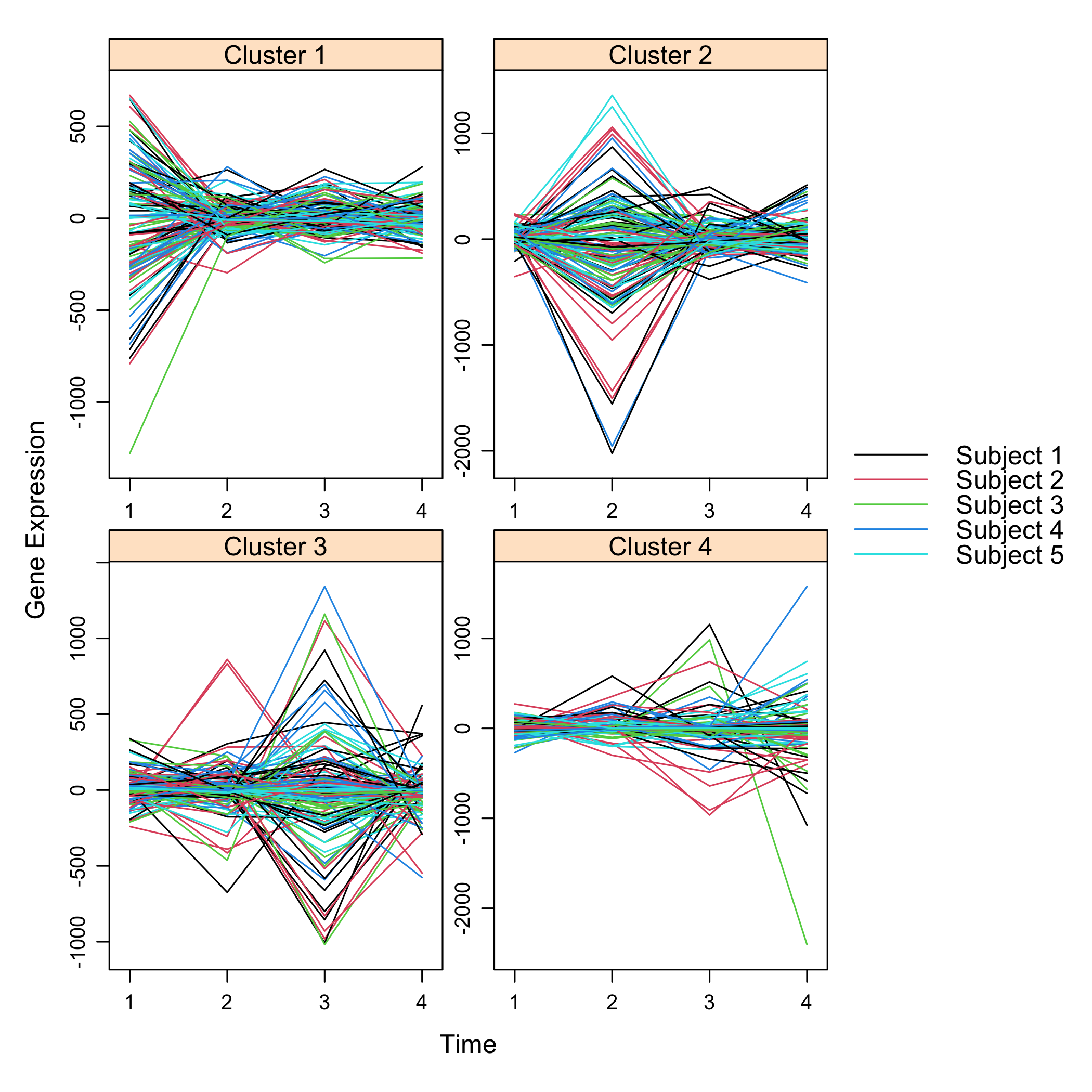
plot of chunk plotsimuldata
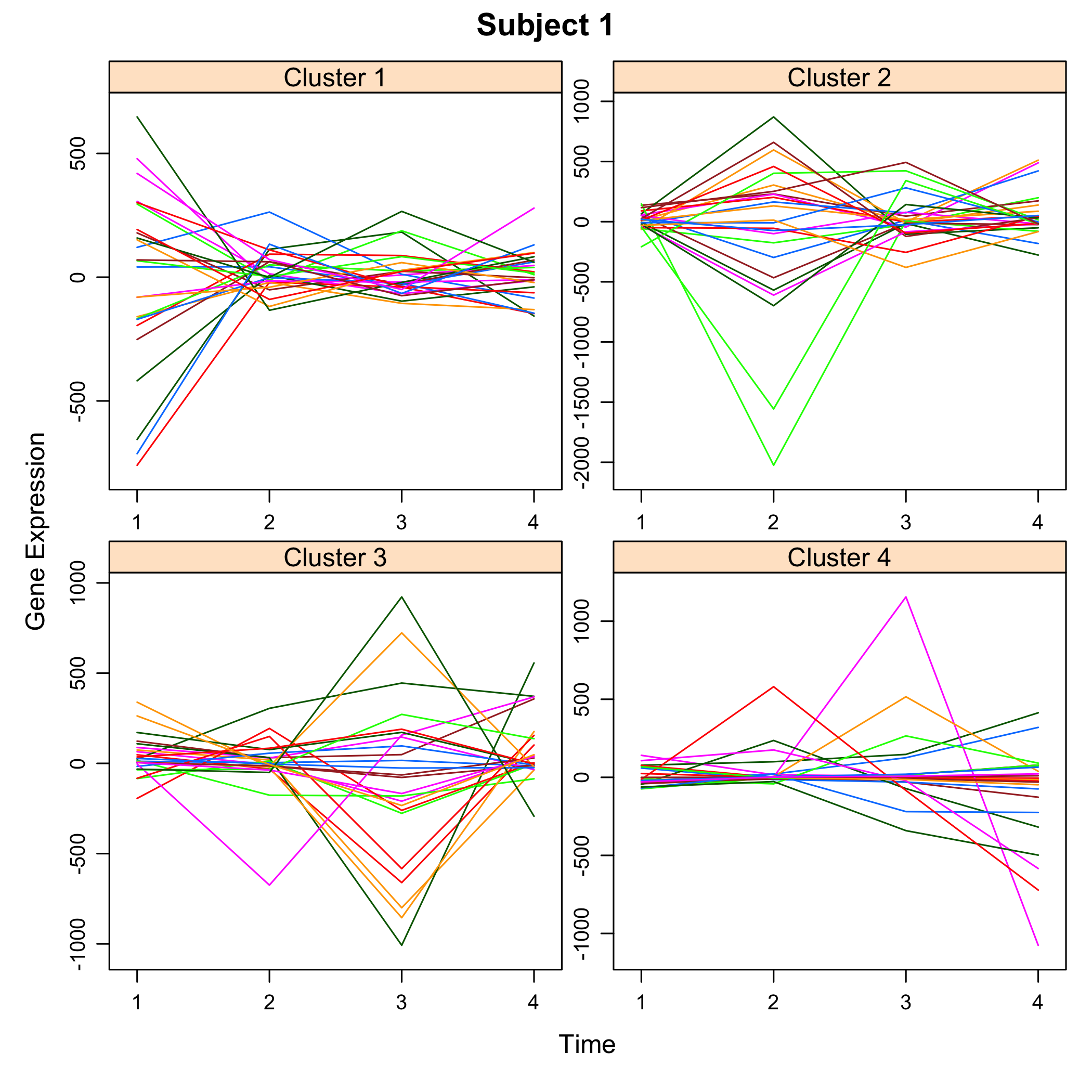
plot of chunk plotsimuldata
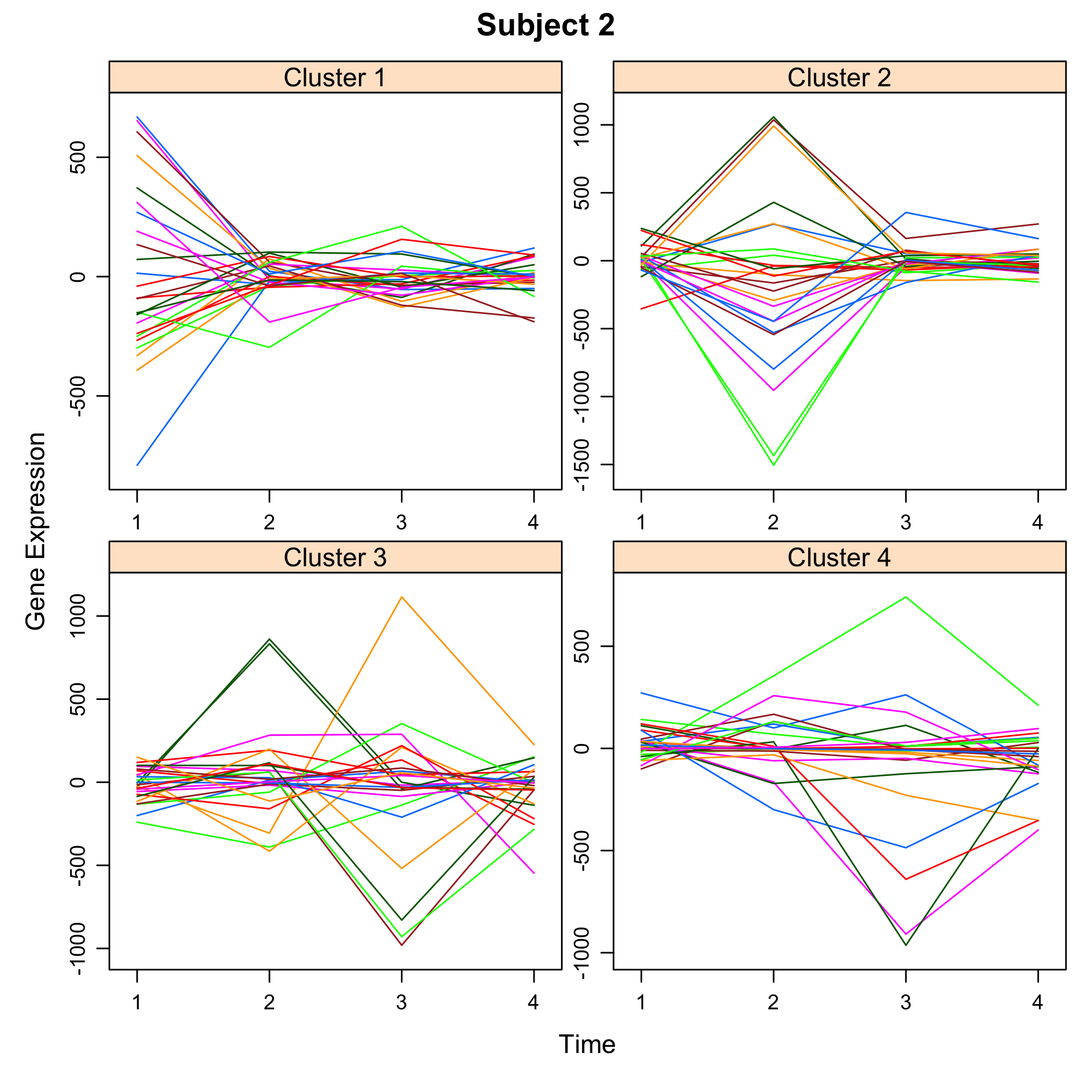
plot of chunk plotsimuldata
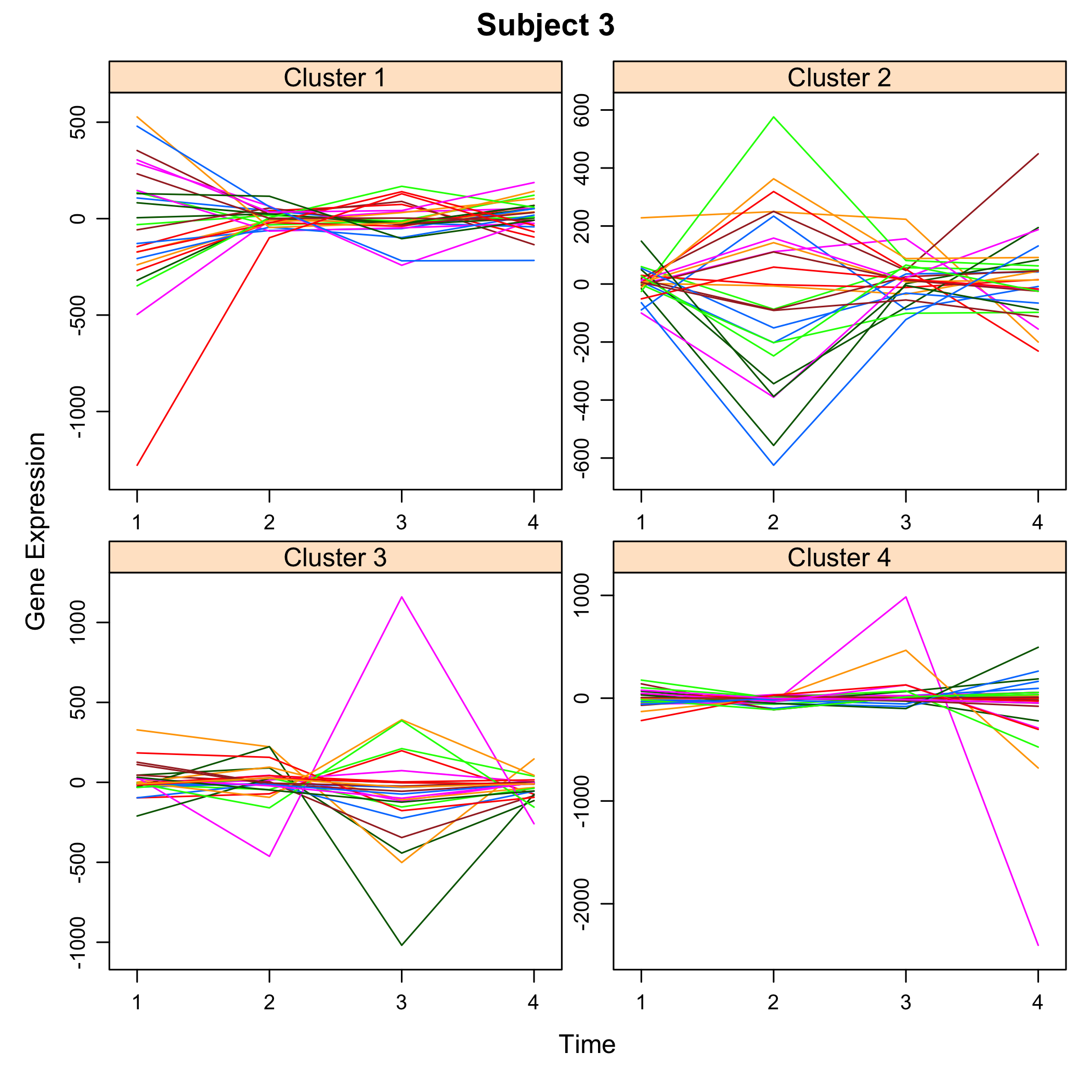
plot of chunk plotsimuldata
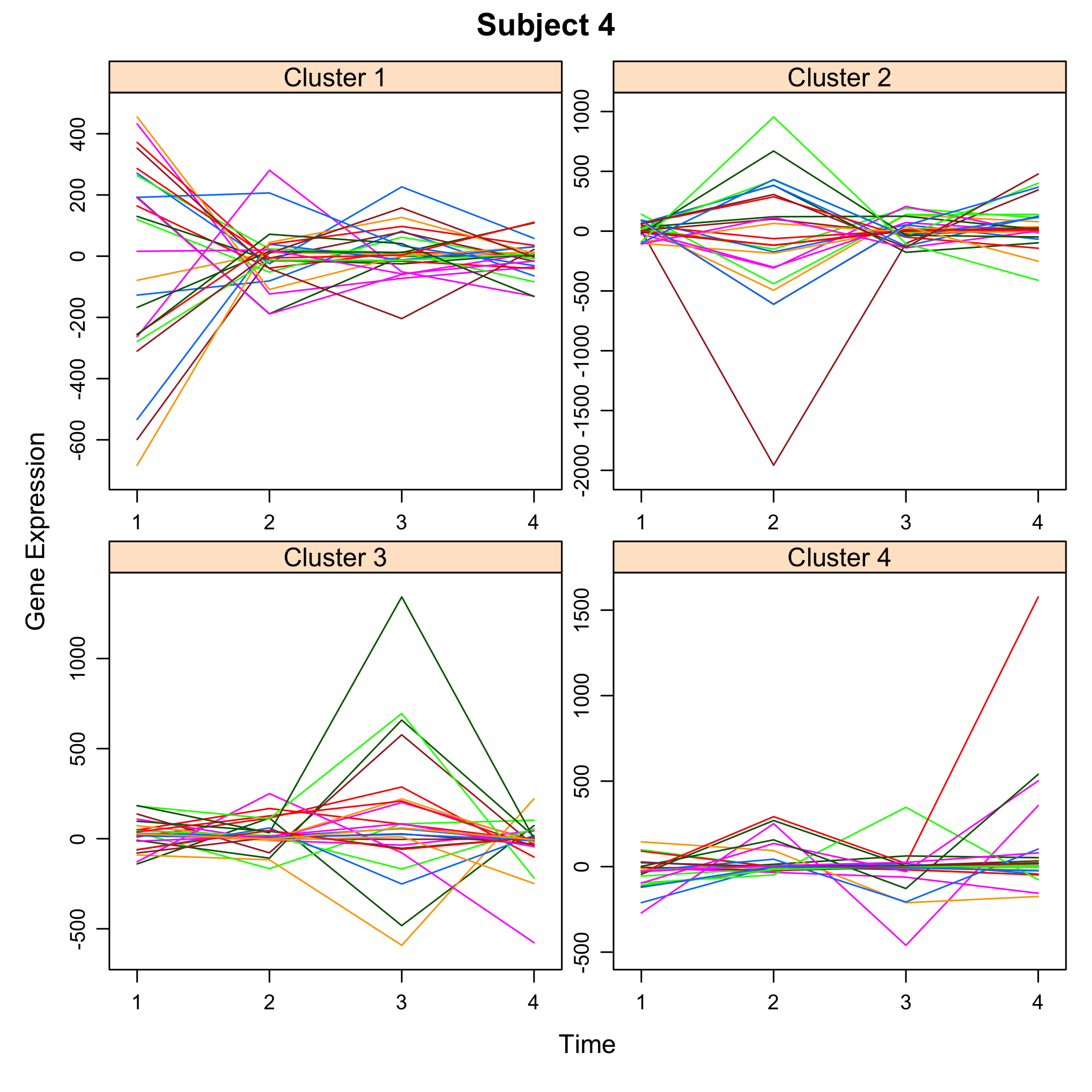
plot of chunk plotsimuldata
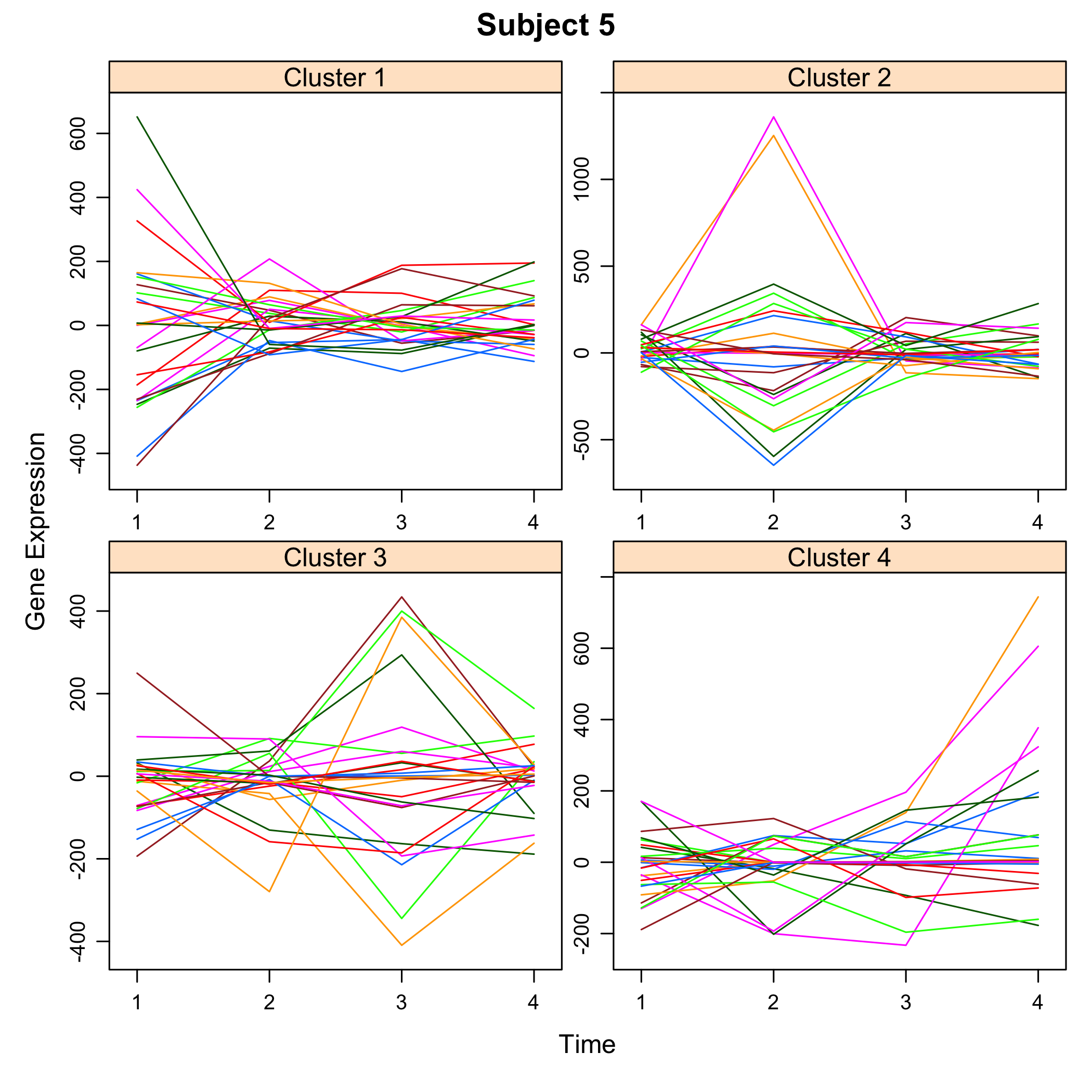
plot of chunk plotsimuldata
Network inference
We infer the new network using subjectwise leave one out cross-validation (all measurement from the same subject are removed from the dataset)
Net_inf_C <- Cascade::inference(M, cv.subjects=TRUE)
#> Loading required package: nnls
#> We are at step : 1
#> The convergence of the network is (L1 norm) : 0.0072
#> We are at step : 2
#> The convergence of the network is (L1 norm) : 0.00139
#> We are at step : 3
#> The convergence of the network is (L1 norm) : 0.00106
#> We are at step : 4
#> The convergence of the network is (L1 norm) : 0.00095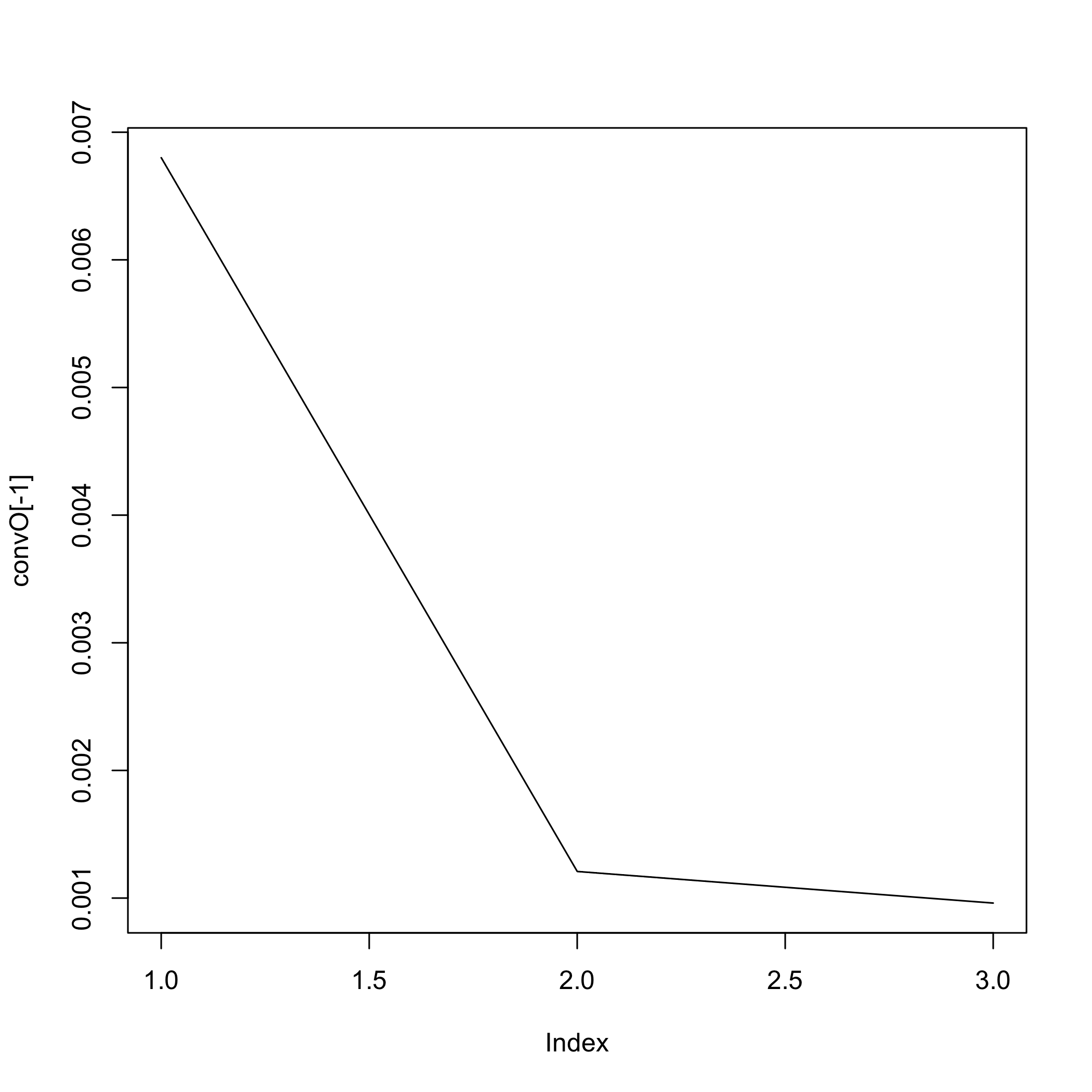
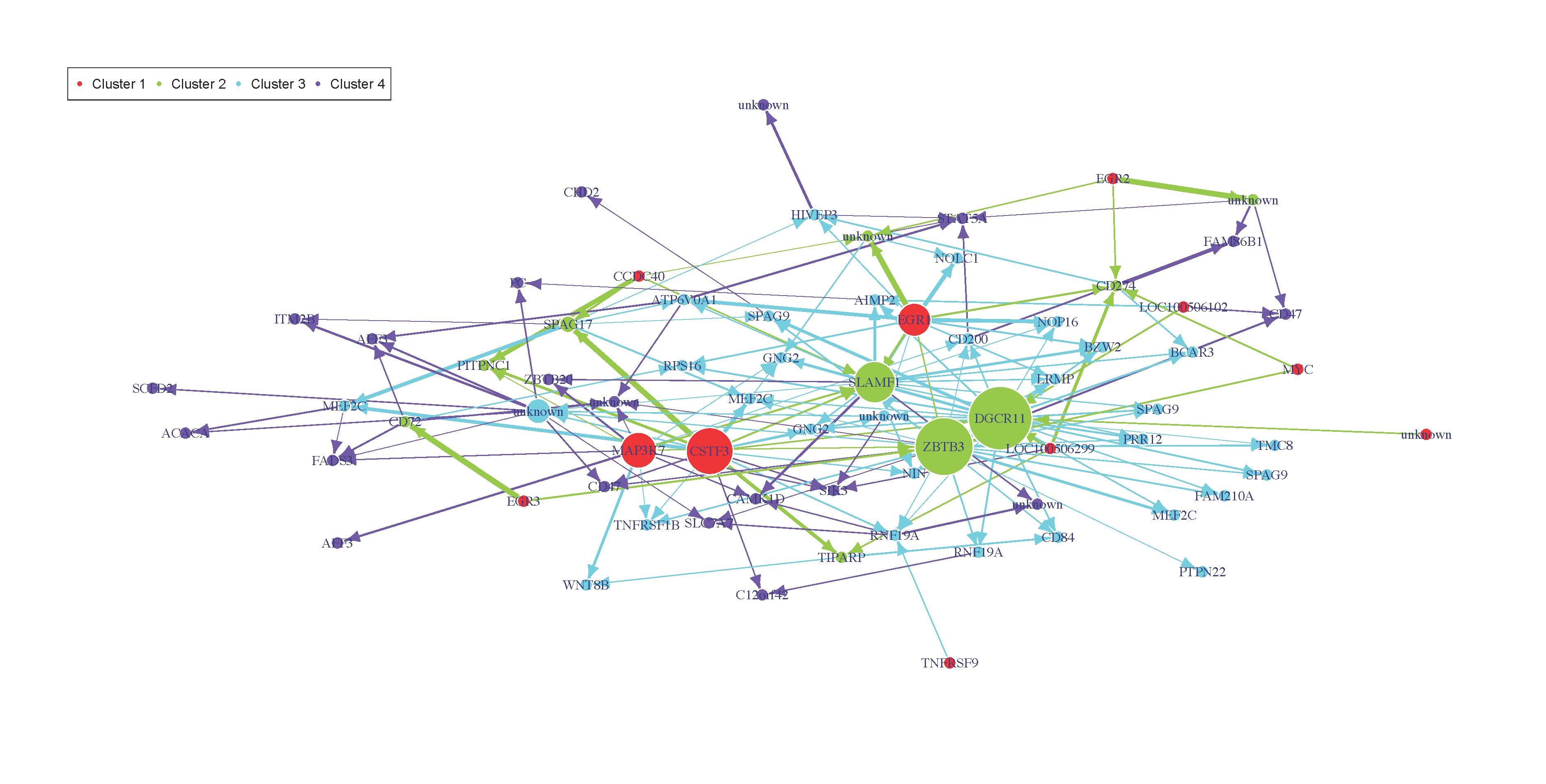
plot of chunk netinf
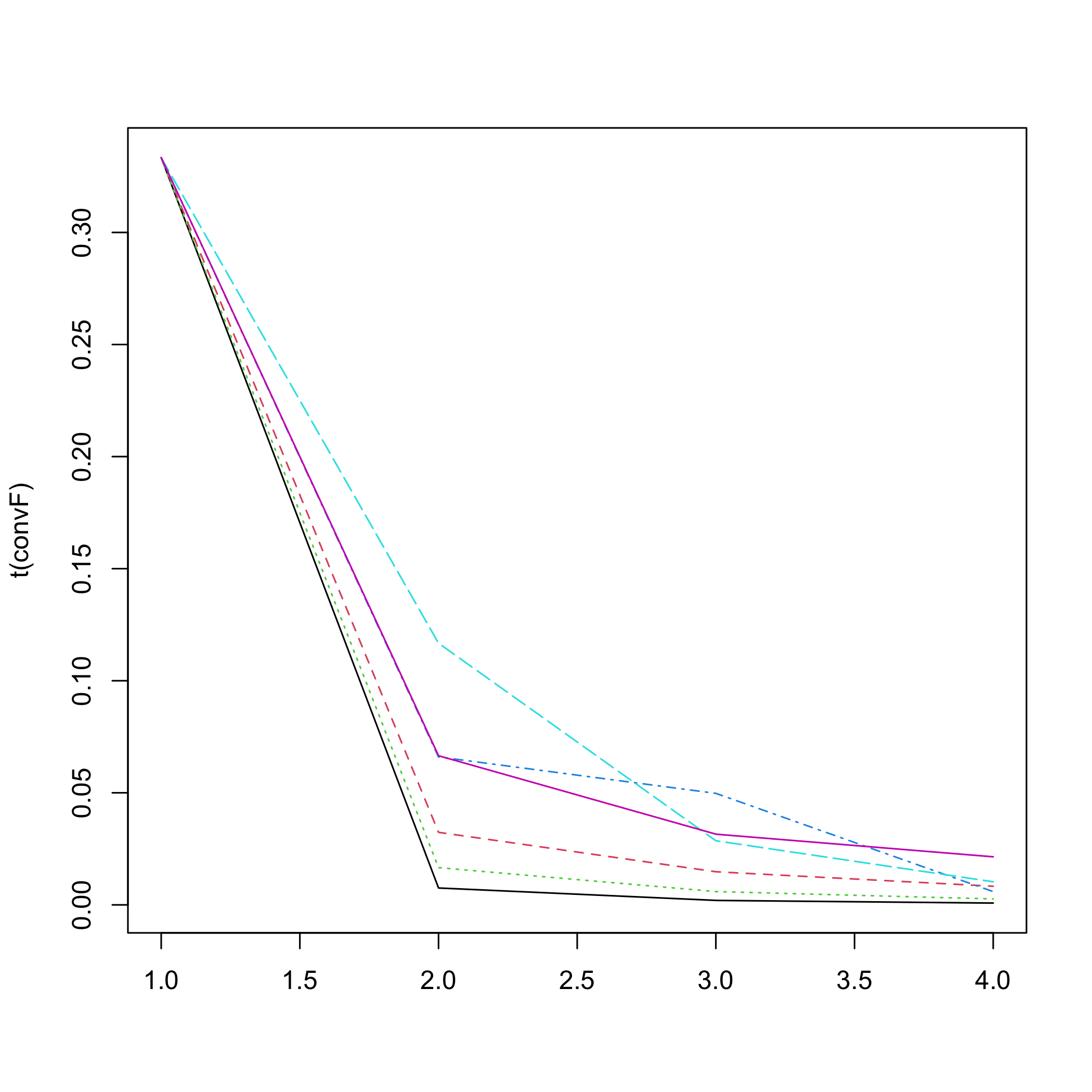
plot of chunk netinf
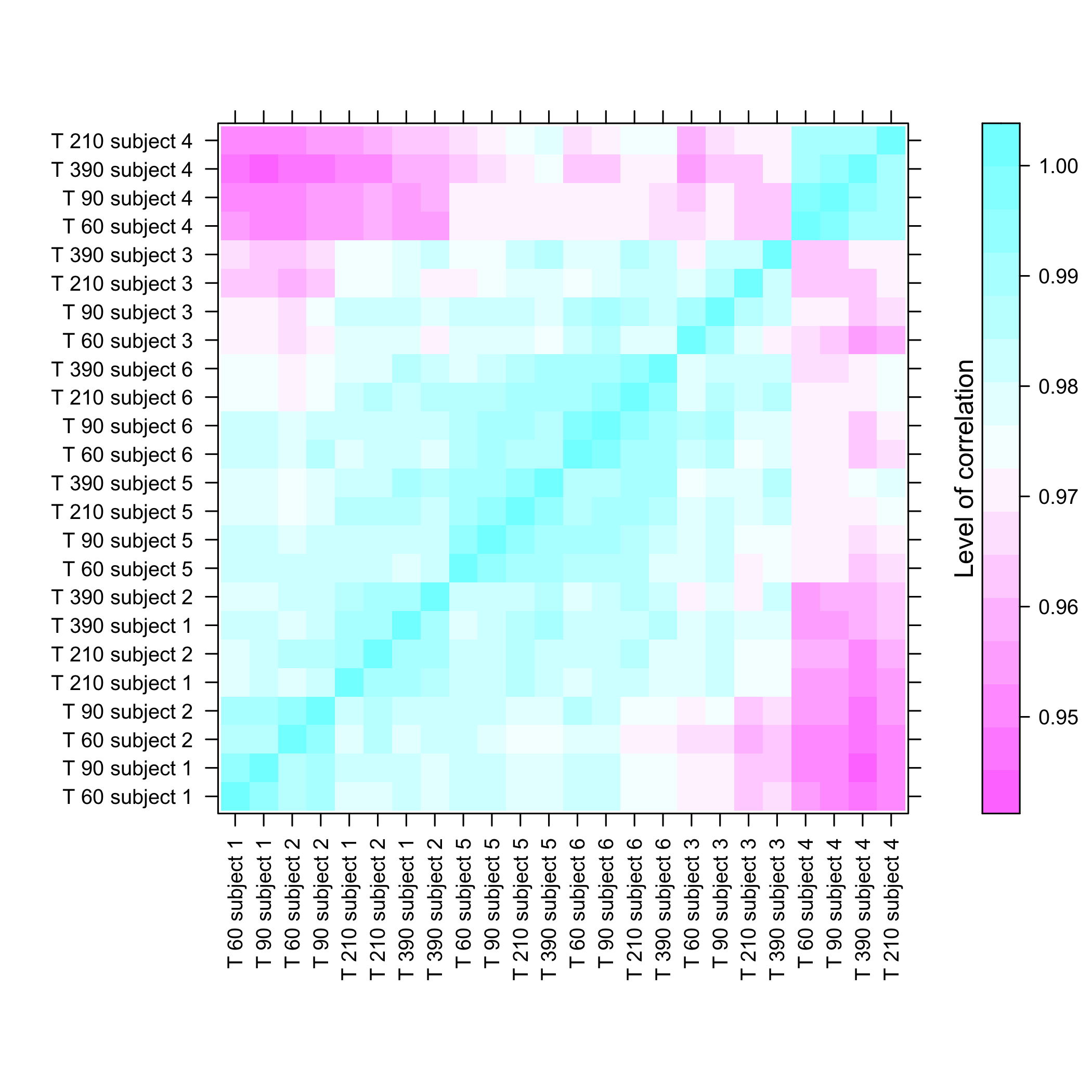
Heatmap of the coefficients of the Omega matrix of the network
stats::heatmap(Net_inf_C@network, Rowv=NA, Colv=NA, scale="none", revC=TRUE)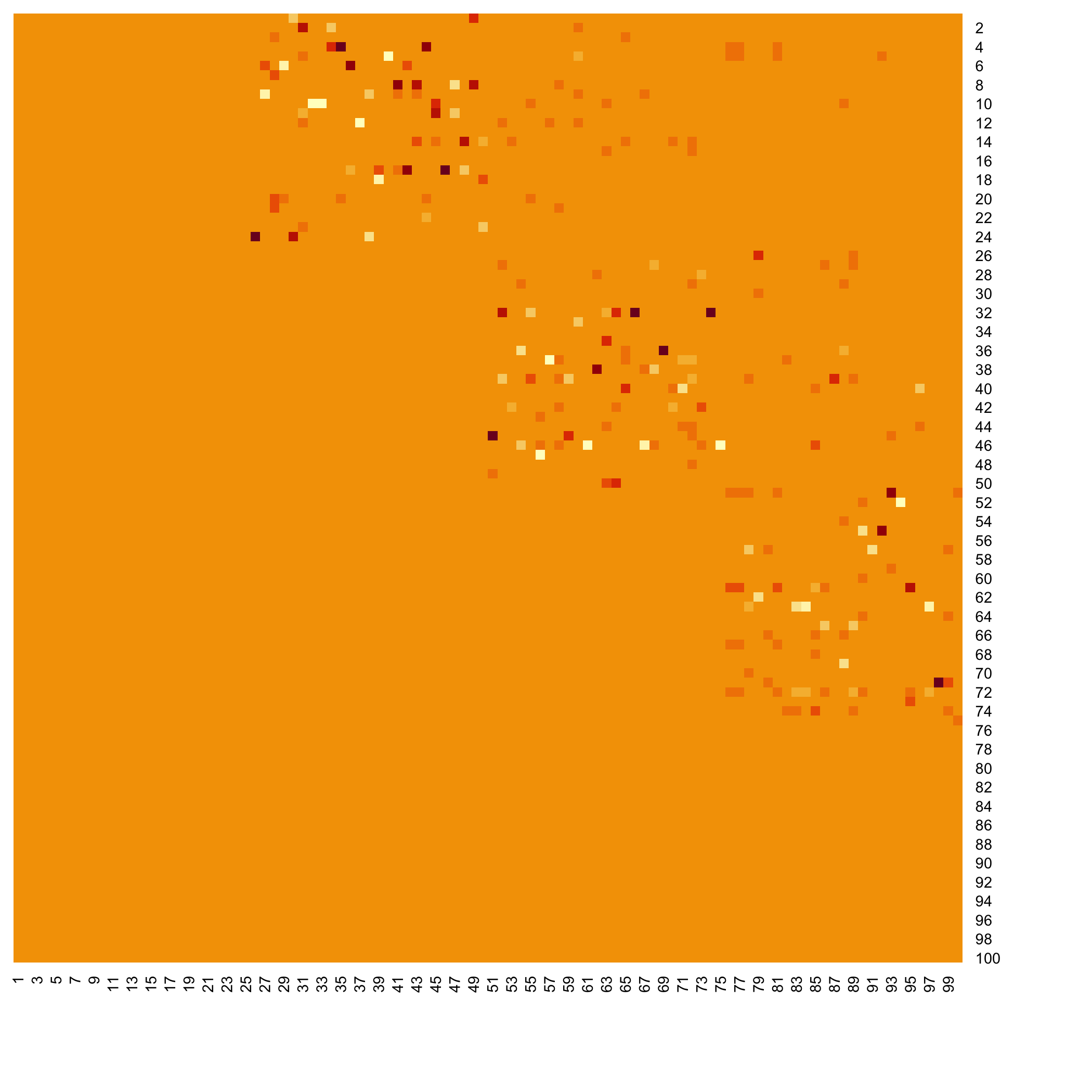
plot of chunk heatresults
###Post inferrence network analysis We switch to data that were derived from the inferrence of a real biological network and try to detect the optimal cutoff value: the best cutoff value for a network to fit a scale free network.
data("network")
set.seed(1)
cutoff(network)
#> [1] "This calculation may be long"
#> [1] "1/10"
#> [1] "2/10"
#> [1] "3/10"
#> [1] "4/10"
#> [1] "5/10"
#> [1] "6/10"
#> [1] "7/10"
#> [1] "8/10"
#> [1] "9/10"
#> [1] "10/10"
#> [1] 0.000 0.001 0.126 0.112 0.091 0.584 0.885 0.677 0.604 0.363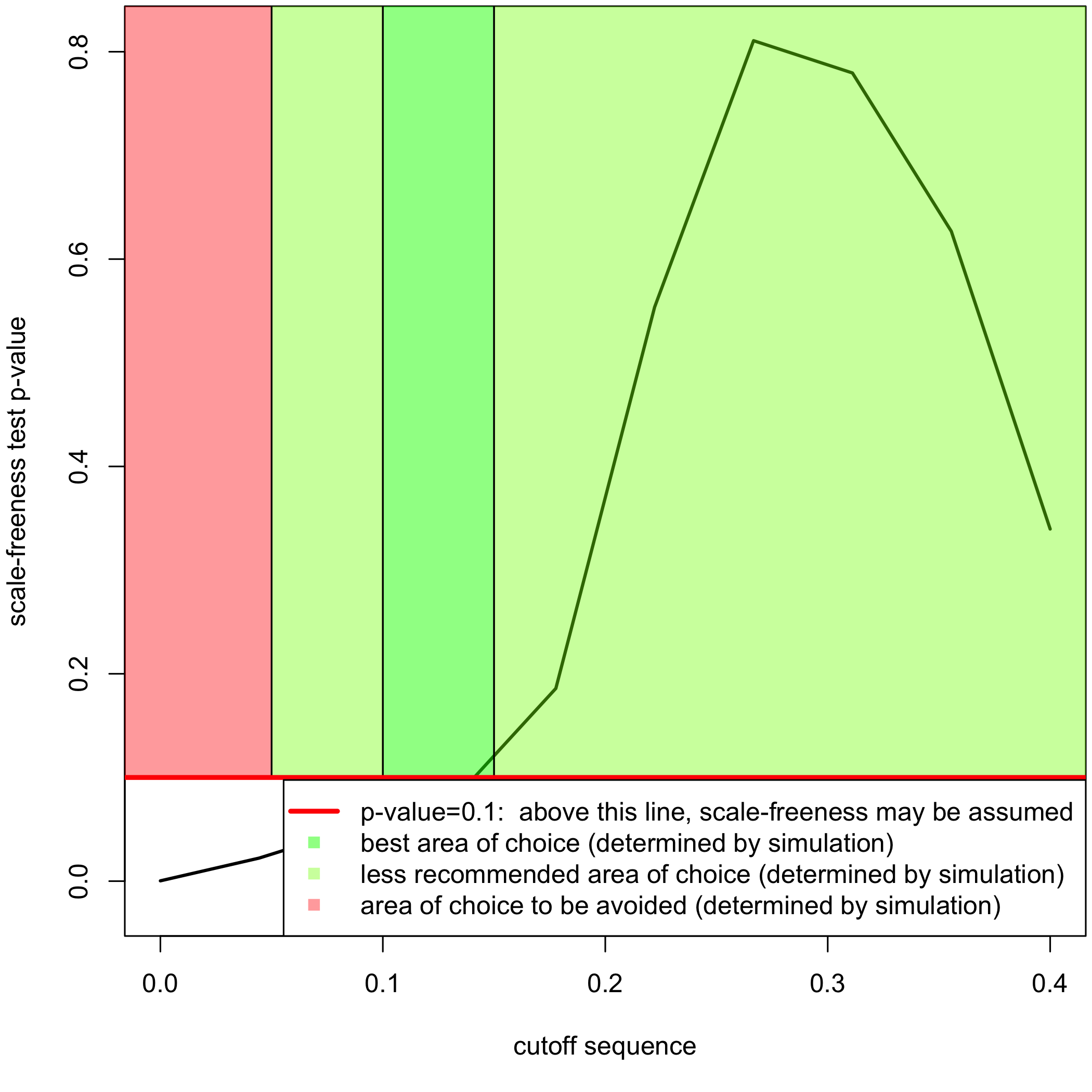
#> $p.value
#> [1] 0.000 0.001 0.126 0.112 0.091 0.584 0.885 0.677 0.604 0.363
#>
#> $p.value.inter
#> [1] 0.0003073808 0.0222769859 0.0521597921 0.0819131661
#> [5] 0.1859011443 0.5539131661 0.8106723719 0.7795175496
#> [9] 0.6267996116 0.3396322205
#>
#> $sequence
#> [1] 0.00000000 0.04444444 0.08888889 0.13333333 0.17777778
#> [6] 0.22222222 0.26666667 0.31111111 0.35555556 0.40000000Analyze the network with a cutoff set to the previouly found 0.14 optimal value.
analyze_network(network,nv=0.14)
#> node betweenness degree output closeness
#> 1 1 0 3 0.8133348 16.4471148
#> 2 2 0 3 0.8884602 7.9547696
#> 3 3 0 1 0.1749376 10.0055952
#> 4 4 0 3 0.5159878 11.4854812
#> 5 5 0 0 0.0000000 0.0000000
#> 6 6 0 13 3.5794097 25.6388630
#> 7 7 0 4 0.9685114 7.0356510
#> 8 8 0 0 0.0000000 0.0000000
#> 9 9 0 0 0.0000000 0.0000000
#> 10 10 3 2 0.6047036 3.0439695
#> 11 11 31 10 1.9146802 8.2263869
#> 12 12 1 1 0.2056836 0.8489352
#> 13 13 97 19 3.7578360 18.6066356
#> 14 14 0 0 0.0000000 0.0000000
#> 15 15 0 0 0.0000000 0.0000000
#> 16 16 0 0 0.0000000 0.0000000
#> 17 17 2 2 0.3985715 1.6450577
#> 18 18 9 1 0.1408025 0.5811461
#> 19 19 0 0 0.0000000 0.0000000
#> 20 20 0 0 0.0000000 0.0000000
#> 21 21 0 0 0.0000000 0.0000000
#> 22 22 0 0 0.0000000 0.0000000
#> 23 23 0 0 0.0000000 0.0000000
#> 24 24 0 0 0.0000000 0.0000000
#> 25 25 0 0 0.0000000 0.0000000
#> 26 26 0 0 0.0000000 0.0000000
#> 27 27 0 0 0.0000000 0.0000000
#> 28 28 9 2 0.3786198 1.5627095
#> 29 29 0 0 0.0000000 0.0000000
#> 30 30 28 6 1.1216028 4.6292854
#> 31 31 0 0 0.0000000 0.0000000
#> 32 32 0 0 0.0000000 0.0000000
#> 33 33 0 0 0.0000000 0.0000000
#> 34 34 3 1 0.1988608 0.8207750
#> 35 35 0 0 0.0000000 0.0000000
#> 36 36 0 0 0.0000000 0.0000000
#> 37 37 0 0 0.0000000 0.0000000
#> 38 38 0 0 0.0000000 0.0000000
#> 39 39 0 0 0.0000000 0.0000000
#> 40 40 0 0 0.0000000 0.0000000
#> 41 41 0 0 0.0000000 0.0000000
#> 42 42 0 0 0.0000000 0.0000000
#> 43 43 0 0 0.0000000 0.0000000
#> 44 44 0 0 0.0000000 0.0000000
#> 45 45 0 0 0.0000000 0.0000000
#> 46 46 0 0 0.0000000 0.0000000
#> 47 47 0 0 0.0000000 0.0000000
#> 48 48 0 0 0.0000000 0.0000000
#> 49 49 0 0 0.0000000 0.0000000
#> 50 50 0 0 0.0000000 0.0000000
#> 51 51 0 0 0.0000000 0.0000000
#> 52 52 0 0 0.0000000 0.0000000
#> 53 53 0 0 0.0000000 0.0000000
#> 54 54 0 0 0.0000000 0.0000000
#> 55 55 0 10 3.2277268 22.2234874
#> 56 56 13 3 0.7360000 3.5063367
#> 57 57 0 0 0.0000000 0.0000000
#> 58 58 1 1 0.2291004 0.9455855
#> 59 59 0 0 0.0000000 0.0000000
#> 60 60 0 2 0.3955933 7.4313822
#> 61 61 0 3 1.2813639 7.0435303
#> 62 62 0 0 0.0000000 0.0000000
#> 63 63 0 0 0.0000000 0.0000000
#> 64 64 2 2 0.3878745 1.6009071
#> 65 65 0 2 1.2169141 10.8093303
#> 66 66 5 1 0.3016614 1.2450723
#> 67 67 3 3 0.5958934 2.4594808
#> 68 68 0 0 0.0000000 0.0000000
#> 69 69 0 0 0.0000000 0.0000000
#> 70 70 0 0 0.0000000 0.0000000
#> 71 71 26 8 1.6479964 6.8019142
#> 72 72 0 0 0.0000000 0.0000000
#> 73 73 0 0 0.0000000 0.0000000
#> 74 74 0 0 0.0000000 0.0000000